Acknowledgements
The authors would like to thank the staff who were involved in the successful delivery of this project from UK Health Security Agency and Agri-Food and Biosciences Institute (AFBI) through the testing of samples and isolates. Large scale surveys are impossible to complete without the work and dedication of the laboratory personnel. We are also grateful to Hallmark Meat Hygiene Ltd. for their collaboration in collecting the whole-head lettuce samples.
AMR Terminology
Internationally, epidemiological cut-off values (ECOFFs) have been used to determine ‘resistance’ (i.e. susceptibility to antimicrobials) in surveillance projects, but it should be noted that ECOFFs do not necessarily indicate clinical resistance (i.e. whether a treatment is likely to be unsuccessful). ECOFFs distinguish between individuals within a species which have or have not developed any phenotypically detectable acquired resistance. The ECOFF is the upper end of the wild type of the species for the agent and does not necessarily indicate clinical resistance. This is because ECOFF determination does not take into account results of clinical efficacy studies, dosing and route of administration of antimicrobial agents, nor the drug’s pharmacokinetic and pharmacodynamic parameters. The Advisory Committee on Microbiological Safety of Food Working Group on Antimicrobial Resistance (ACMSF AMR WG) has published recommendations on the use of AMR terminology/nomenclature used in FSA reports which includes how best to integrate ECOFFs, clinical breakpoint and genomic data to most accurately reflect the AMR status of bacterial isolates found in food surveys. Results in this study are based on ECOFFs.
Lay Summary
Whole-head lettuce is a widely consumed food in the UK. As part of surveillance of microbiological risks in the food chain, it is important to monitor the presence of microbial pathogens and antimicrobial resistant (AMR) bacteria in fresh produce, as it is often consumed raw. Moreover, consumption of lettuce has been associated with outbreaks of foodborne illness. This survey was carried out to generate new data on the presence of Shiga-toxin producing E. coli (STEC), Salmonella, E. coli and levels of AMR found in these organisms in whole-head lettuce on retail sale in the UK.
A total of 313 whole-head lettuce samples were collected from major supermarket chains within the UK between October 2024 and September 2025. Samples were tested for the presence of bacteria that can make people ill when present in lettuce (STEC and Salmonella) and bacteria that indicate hygiene issues during the production or handling of the food before reaching the consumer (E. coli). When any of these bacteria were found in the whole-head lettuce samples, we did further testing to ascertain whether they carried antimicrobial resistance, which can make treatment of infections more difficult. The main results of this survey are:
-
No STEC, Salmonella and AMR bacteria were detected from any of the whole-head lettuce samples tested.
-
E. coli was detected in 7.7% (24 out of 313) of whole-head lettuce samples, albeit at low levels (all samples apart from one had a level of less than 20 CFU/g).
-
E. coli was more common in whole-head lettuce samples tested during the summer (June to August 2025).
The data presented in this report provides reassurance to consumers and regulators that the likelihood of being exposed to AMR bacteria or acquiring a foodborne illness caused by STEC or Salmonella through consumption of whole-head lettuce at retail sale in major UK supermarkets is low.
Executive Summary
Monitoring microbiological hazards in the food chain is required to ensure that food reaching consumers is safe. As well as the presence of pathogenic bacteria (that make people ill), it is important to survey antimicrobial resistance (AMR). AMR is a major public health concern and infections in humans caused by bacteria resistant to antibiotics are associated with substantial morbidity and mortality. Lettuce is widely consumed raw in the UK and has been associated to outbreaks of foodborne illness. The risk of exposure to consumers of any AMR bacteria present in lettuce would be higher compared to other foods that are consumed after cooking/heating.
This study assessed the prevalence of microbiological and AMR hazards in samples of whole-head lettuce collected at major retail supermarkets in the UK. Between October 2024 and September 2025, 313 samples were collected. Samples were tested for Shiga-toxin producing E. coli (STEC) and Salmonella and for E. coli including those with extended-spectrum beta lactamase (ESBL) and/or AmpC-phenotypes or with resistance to carbapenems and colistin. All E. coli isolates obtained were also tested to determine the Minimum Inhibitory Concentrations (MIC) for a range of antimicrobials.
No STEC or Salmonella were isolated from any of the samples, but one sample had traces of stx DNA (gene encoding for the Shiga toxin). E. coli was detected in 24 samples (7.7%) with the majority with levels of <20 CFU/g (i.e. only one sample could be enumerated at 20 CFU/g). None of the E. coli isolates showed any resistance to any of the antibiotics tested. No ESBL/AmpC-producing E. coli, colistin resistant E. coli or carbapenemase-producing E. coli were detected in any of the whole-head lettuce samples tested. Whole-head lettuce samples collected during the summer months (June to August 2025) were significantly more likely to test positive for E. coli.
In conclusion, no STEC, Salmonella or AMR E. coli were detected in this survey, with a small proportion of samples having low levels of E. coli. These results confirm a low prevalence and low risk to UK consumers of whole-head lettuce in relation to the microbiological hazards investigated.
Introduction
Antimicrobials are the cornerstone of modern human and veterinary medicine and are also used in agricultural practices. Microorganisms becoming resistant to antimicrobials represents a major public health threat. The UK is currently working on the second 5-year National Action Plan (NAP), “Confronting antimicrobial resistance 2024 to 2029” (Department of Health and Social Care (DoHSC), 2024) with the aim of controlling and containing antimicrobial resistance (AMR) by 2040. Taking a One Health approach across people, animals, food and the environment is essential to implement effective interventions that reduce the burden of antimicrobial resistance. One key theme in the current plan is reducing the unintentional exposure to antimicrobials, with strengthening surveillance in the environment and the food chain being an important aspect to achieve this goal.
In the UK, surveillance studies to monitor AMR in the food chain have primarily focused on meat at retail sale (Food Standards Agency (FSA), 2021, 2022). However, the scope of AMR surveillance has been broadened following a review of the Food Standards Agency’s AMR programme conducted by independent experts in 2023. While continued surveillance of poultry meat was still considered a priority, fresh produce was also identified as important for surveillance programmes and was ranked second on the priority list (Nicholls et al., 2023). This was primarily driven by challenges in current agricultural practices, with multiple opportunities for introduction of foodborne pathogens, and an understanding that fresh produce is often eaten raw after minimal processing, without a “kill” step to reduce microbiological load (Thomas et al., 2024), which would also increase the risk of exposure to any AMR bacteria in this food commodity (Nicholls et al., 2023).
E. coli are commensal bacteria mainly found in the gut of warm-blooded animals but are also a leading cause of infection (Foster-Nyarko & Pallen, 2022). The presence of E. coli in food can be indicative of faecal contamination, poor hygiene, cross-contamination or other production issues and, while often not representing a direct risk to health, E. coli bacteria can act as a useful marker to identify conditions leading to an increased risk of contamination with human pathogens (UK Health Security Agency (UKHSA), 2024). Due to their wide distribution and importance, E. coli are part of established human and livestock AMR surveillance programmes and also make a useful indicator for AMR in the food chain across different food commodities (Anjum et al., 2021; Nicholls et al., 2023).
Lettuce is a widely consumed food commodity that is almost always eaten raw. Consumption of lettuce has been associated with outbreaks of foodborne illness. For example, in 2019 an outbreak with 32 cases of STEC O26:H11 in the UK was epidemiologically linked to consumption of pre-packed sandwiches containing a mixed salad of Apollo, iceberg lettuce and spinach leaves (Butt et al., 2021). Similarly, in 2016, 165 cases of STEC O157:H7 occurred in the UK with illness being associated with consumption of mixed salad leaves from a particular supplier (Gobin et al., 2018). STEC outbreaks have also been linked to lettuce in other parts of Europe (Friesema et al., 2008), Japan (Terajima et al., 2014) and the USA (Slayton et al., 2013). Several outbreaks of Salmonella have also been linked to consumption of lettuce. In the UK, an outbreak of 361 cases of S. Typhimurium in England and Wales in the summer of 2000 was linked to consumption of lettuce outside of the home environment (Horby et al., 2003). Similarly, consuming lettuce in a meal outside the home was associated with S. Newport infection in an outbreak affecting 113 cases in Northern Ireland in 2004 (Irvine et al., 2009). In another incident, iceberg lettuce was identified as the common vehicle for multiple outbreaks of S. Braenderup in the West Midlands (Gajraj et al., 2012). Salmonella enterica outbreaks linked to lettuce have also been reported in other parts of the world, including a recent outbreak of S. Typhimurium in day care centres in Finland that affected over 700 people, mostly children, which was epidemiologically and microbiologically linked to a vegetable mix where pre-chopped iceberg lettuce was the most likely culprit (Lehti et al., 2024).
This study aimed to provide data on the prevalence and antimicrobial resistance profiles of STEC, Salmonella and Escherichia coli present in whole-head lettuce available at retail in supermarkets in the UK, to assess public health risks posed by this food commodity to consumers.
Materials and Methods
2.1. Sample collection at retail and transportation to the testing laboratory
Sampling design and collection were completed by HallMark Meat Hygiene Ltd, trading as HallMark Veterinary & Compliance Services. The sampling plan aimed for 315 samples, including a 5% contingency above the recommended 300 samples to account for potential missing or rejected samples. The sampling was conducted over 12 months from October 2024 to September 2025, incorporating seasonal fluctuations in lettuce consumption.
HallMark sampling officers were instructed to collect whole head pre-packaged lettuces either in fully sealed or in partially open-end packaging. These included cultivated lettuces in which the leaves form a dense rosette either with a closed or loose head structure, such as iceberg, romaine, butterhead, little gem, sweet gem and read leaf varieties (Figure 1). Sampling officers followed consumer-like purchasing behaviours when selecting lettuce heads, choosing from different brands and varieties where available. Sample metadata, including product and retailer details, batch/lot numbers and farm origin (where available) were recorded. Bagged salads, composite salads, salad bowls and loose or cut lettuce were excluded from this study. Bagged salad varieties are excluded as these undergo additional processing steps (cutting, washing and bagging) which may impact contamination present. Also bagged salad can consist of several salad varieties (e.g. a composite food) which makes it more difficult to identify the source of contamination if detected.
Sampling was conducted in the top twelve UK supermarkets proportionally selected based on market share data from the Defra Family Food Surveys (2019/20 to 2021/22). A stratified sampling approach based on UK International Territorial Levels (ITL1) regions (East of England, East Midlands, West Midlands, London, North East, North West, South East, South West, Yorkshire and the Humber, Northern Ireland, Scotland and Wales) was used to ensure geographic representativeness and diversity of produce origin, capturing both locally sourced and imported lettuces available at retail across the UK.
This study design ensured that the sampling process was statistically robust, geographically representative, and aligned with FSA surveillance objectives. Minor deviations in regional and retailer allocation from the original plan were mitigated through built-in contingency.
After collection, samples were packed into cold boxes with sufficient ice packs to achieve and maintain a target temperature of <8°C during transit. Samples were dispatched by overnight courier to one of four designated microbiology laboratories: UKHSA Porton FW&E Laboratory, UKHSA London FW&E Laboratory, UKHSA York FW&E Laboratory, or Agri-Food and Biosciences Institute (AFBI), Belfast. While most samples maintained a temperature of <8°C during transit, 13% of samples were received at > 8°C, with the highest temperature being 16.8°C. These samples were included in the testing.
In total, 316 samples were collected, of which 313 samples were classified as satisfactory at the time of receipt at the laboratories, while 3 samples were rejected because they had exceeded their expiry dates. Given the 5% contingency allowance, the total number of satisfactorily tested samples met the project objectives.
Samples were either tested immediately or stored at 2-8°C and tested within 36 hours of receipt, and within the shelf-life of the product.
2.2. Microbiological examination of lettuce samples
Samples were processed by cutting a 40±5g cross-sectional lettuce wedge (figure 2) running the length of the lettuce and incorporating all layers (external, middle and internal leaves and base) on a sterile surface with an autoclaved knife, and preparing a 10-1 homogenate in Buffered Peptone Water (BPW; prepared in-house), according to ISO 6887-1:2017 (International Organization for Standardization (ISO), 2017). A subsection of samples (Terajima et al., 2014) were additionally processed using a “rinse” methodology for comparison. To do this, the remaining sample after cutting two cross-sectional lettuce wedges as indicated above (minimum portion of 100 g) was rinsed with 100 mL of BPW and massaged manually for 2 minutes.
2.2.1. Examination of lettuce samples for the presence of generic and antimicrobial resistant Escherichia coli
A portion (20 mL) of the homogenate in BPW for wedge samples, and rinse samples where applicable, was used to enumerate E. coli using a surface inoculation method or a pour plate method with a tryptone bile X-glucuronide (TBX) agar (prepared in-house or E&O laboratories) based on ISO 16649-2:2001 (International Organization for Standardization (ISO), n.d.).
The remaining ~380 ml of homogenate was incubated at 37°C for 18 h and then sub-cultured on TBX agar (prepared in-house; for the detection of generic E. coli), MacConkey containing 1 mg/L cefotaxime (prepared in-house; for the detection of presumptive ESBL-/AmpC-producing E. coli), MacConkey Agar containing 2 mg/L colistin (prepared in-house; for the detection of colistin-resistant E. coli) and CHROMID® Carba Smart agar plates (Biomerieux; for the detection of carbapenemase-producing E. coli). All plates were incubated at 44 °C for 20± 2 h except the CHROMID® Carba Smart agar plates which were incubated at 37 °C for 22± 2 h. For the rinse samples, 25 mL of the rinse was diluted 1 in 10 in BPW, incubated at 37°C for 18 h and then sub-cultured on TBX agar with plates being incubated at 44°C for 22± 2 h. Isolate identity was confirmed as E. coli using matrix assisted laser desorption ionization MALDI-TOF (Bruker) MS or biochemical testing.
2.2.2. Examination of lettuce samples for the presence of Salmonella species and Shiga-toxin producing Escherichia coli
For the detection of Salmonella and STEC, methods based on ISO 6579:1:2017 (International Organization for Standardization (ISO), 2020) and ISO/TS 13136:2012 (International Organization for Standardization (ISO), 2012) respectively were followed. After incubation (37°C for 18 h) of the homogenate (and diluted rinse where applicable), a portion was heat inactivated and analysed by real-time PCR using the SureTect™ Salmonella species, Typhimurium and Enteritidis Multiplex PCR kit (Thermofisher Scientific) and the SureTect™ Escherichia coli O157:H7 and STEC Screening PCR Assay (Thermofisher Scientific) following the manufacturer’s instructions.
2.3. Determination of minimum inhibitory concentrations for E. coli isolates
One isolate from each isolation medium for each sample was selected to determine the minimum inhibitory concentrations (MIC) for a range of antimicrobials in accordance with Assimilated Decision 2020/1729/EU. A broth microdilution method was used to determine the MICs using SensititreTM broth microdilution (BMD) plates. Isolates were inoculated into Mueller Hinton broth at a suitable concentration and dispensed onto commercially prepared plates (Sensititre EUVSEC3, Thermofisher) containing two-fold dilution series of antimicrobials in accordance with Assimilated Decision 2020/1729/EU. After incubation at 35°C for 18-24 h the plates were examined and growth end-points were established for each antimicrobial to provide MICs. Microbiologically-resistant and susceptible interpretations for the MICs were obtained by comparison with epidemiological cut-off values (ECOFFs) specified in EU Decision 2020/1729 (European Commission (EU), 2020). ECOFF values separate the naive, susceptible bacterial populations from isolates that have developed reduced susceptibility to a given antimicrobial agent, as recommended in the European Centre for Disease Prevention and Control (ECDC) EU protocol for harmonised monitoring of antimicrobial resistance in human Salmonella spp. and Campylobacter spp. Isolates (European Centre for Disease Prevention and Control (ECDC), 2024). The ECOFFs differ from breakpoints used for clinical purposes, which are defined against a background of clinically relevant data.
2.4. Statistical analysis and data visualisation
Fisher’s exact test was used to compare sample groups where relevant using Microsoft Excel 97-2003 and an on-line calculator (Analyze a 2x2 contingency table) with a significance level set at 0.05. Linear regression analysis and maps were created using RStudio 2024.04.2.
2.5. Quality assurance
All the laboratories which have performed testing in this study are UKAS-accredited to ISO 17025 and participate in External Quality Assurance (EQA) schemes. All analyses were performed by trained and competent staff.
Results
3.1. Description of samples
A total of 313 samples of different varieties of whole-head lettuce (table 1) were collected and tested between October 2024 and September 2025. Samples were purchased from 191 different stores/shops belonging to the top 12 largest supermarket chains (table 2). Only 12% of lettuce heads collected were labelled as organic (table 3). Country of origin was recorded for all samples from the product label (table 4). Most samples originated from United Kingdom (51%) followed by Spain (47%), but the distribution of home-grown and imported lettuce varied during the year. While 95% of samples originated from UK in summer, only 15% of samples collected during the winter months were UK grown. During autumn and spring 47% and 39% of samples originated from UK, respectively (table 5). Due to lack of clear labelling or traceability information on the product packaging, the farm of origin could not be determined for 36% of samples (n=113).
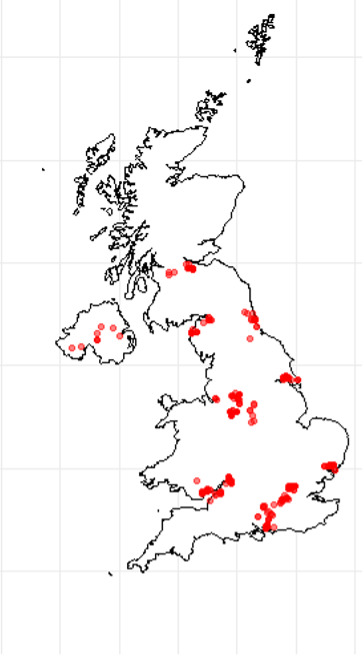
Samples were collected across all regions of England (East, East Midlands, London, Northeast, Northwest, Southeast, Southwest, West Midlands, and Yorkshire and the Humber), Scotland, Wales and Northern Ireland. Sampling locations are illustrated in figure 3.
3.2. Microbiological results
Salmonella and STEC were not isolated in any of the 313 samples (table 6). There was a weak PCR signal for stx DNA in one sample (Ct = 38.3), but STEC was not isolated.
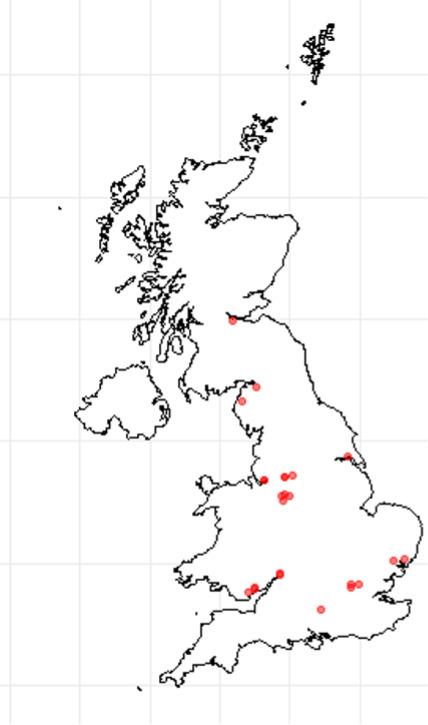
β-glucuronidase positive E. coli were detected in 24 out of 313 (7.7%) samples processed by cutting a wedge of the whole head lettuce (table 6). Samples where E. coli was detected were purchased across the UK (figure 4). Of those samples, the level of E. coli was less than 20 CFU/g in 23 samples and 20 CFU/g in one sample (an iceberg lettuce from the UK). Specific AMR E. coli targeted through selective culturing (i.e. ESBL-/AmpC-producing, carbapenem-resistant and colistin-resistant E. coli) were not detected in any samples (Table 6).
A subset of 12 samples were additionally tested using a rinse of the remaining lettuce, to compare both methodologies (table 7). No STEC or Salmonella were detected in these samples by either methodology. E. coli was detected in three samples using both processing methods and we did not detect E. coli with either methodology in seven samples. In two samples E. coli was detected when processing from a wedge of the lettuce but not when processing from the rinse. The enumeration results, while not directly comparable between the two methodologies, were consistent between the methods and only two samples overall had a level of E. coli above the limit of detection of the enumeration assays. One was an iceberg lettuce sample from the UK (20 CFU/g when processing a wedge of lettuce), and the other was a red leaf lettuce from the UK (200 CFU/ portion rinsed using the rinse methodology). There was no statistically significant difference in E. coli detection rates between the two methods (Fisher’s exact test; p= 0.6). In total, 27 E. coli isolates (24 isolated from lettuce wedges and 3 isolated from lettuce rinses) were analysed using broth microdilution to determine their MIC. All 27 isolates were fully susceptible to all antibiotics tested (Table 6).
The summer period was strongly associated with increased detection rates of E. coli (Fisher’s exact test; p=0.0013; table 5). UK was recorded as the country of origin for 75% of samples with E. coli, with the other 25% of positive samples originating from UK. The difference in E. coli detection frequency between samples originating in UK (18/160; 11%) and outside of UK (6/153; 4%) was statistically significant (Fisher’s exact test; p = 0.0185). However, after adjusting for season, lettuce grown in the UK or outside of the UK did not differ significantly in E. coli detection rates. No significant differences were found when considering the type of lettuce (table 1), the supermarket of purchase (table 2) or whether the samples were labelled as organic (table 3).
Discussion
Whole-head lettuce samples collected from October 2024 to September 2025 from supermarkets in the UK demonstrated good microbiological quality with regards to all parameters tested. No Salmonella or STEC were isolated from any samples. E. coli was detected in 7.7% of samples but the levels of contamination were low, with all samples being considered “satisfactory” (<20 CFU/g) except for one sample (20 CFU/g) that would be classified as “borderline” according to the UKHSA guidelines for assessing the microbiological safety of ready-to-eat foods placed on the market (UK Health Security Agency (UKHSA), 2024). Moreover, no AMR E. coli were detected in any of the lettuce samples tested in this survey.
Throughout the survey, we processed samples by cutting a cross-sectional wedge including all layers of lettuce (external, middle and internal leaves) and the corresponding part of the base. The outer leaves and base might have a greater chance to become contaminated from bacteria present in soil, irrigation water or animal faeces, which have been linked to bacterial outbreaks of foodborne illness associated with fresh produce (Thomas et al., 2024). However, the inner, younger leaves of romaine lettuce have been shown to produce exudates rich in nitrogen and carbon and to support growth of E. coli O157:H7 to significantly higher population sizes compared to middle leaves (Brandl & Amundson, 2008). Therefore, any bacteria reaching inner leaves might have a greater availability of nutrients to grow under the right conditions and it was decided that a cross-sectional wedge would accurately represent all parts of the whole-head lettuce with a potential to contribute to microbial contamination. To provide further assurance of the suitability of the method, a fraction of samples was processed in parallel by rinsing the whole-head lettuce with comparable results.
The prevalence of E. coli in lettuce and salad leaves has been investigated in previous studies in the UK. The Food, Water and Environment Microbiology Service Laboratory (FWEMS) at UK Health Security Agency (UKHSA; previously Public Health England) has conducted two recent UK surveys of salad products collected from retail and catering businesses during 2023-24 and 2020-21 (Kesby et al., 2026; McLauchlin et al., 2022). For leafy salad ingredients, the first survey conducted in 2020-21 reported 13 (6.5%) out of 201 of samples with a level of E. coli of >20 CFU/g (McLauchlin et al., 2022) while the more recent survey conducted in 2023-24 reported 15 (3.5%) out of 432 samples of leafy salad ingredients with that level of E. coli (Kesby et al., 2026). In the current survey, only 0.3% of samples (n=1) showed an E. coli level of ≥ 20 CFU/g. Different sample types and premises were included in these studies, which might explain the difference in the E. coli levels observed. While the current study only included whole-head lettuce samples from supermarket premises, samples in previous surveys were collected from supermarkets, catering premises and other retailers (e.g. independents), and included both loose and prepacked bagged products. It is possible that cross-contamination during storage, processing or handling of the lettuce is responsible for the elevated E. coli results in previous surveys compared to the current survey. Of note, McLauchlin (McLauchlin et al., 2022) reported 15% ready-to-eat salad products with E. coli results ≥ 20 CFU/g from catering premises compared to 1% from major national supermarket retailers.
Although E. coli detection rates and levels were low in this survey, the proportion of whole-head lettuce samples with E. coli was significantly higher in the summer. Similarly, McLauchlin (McLauchlin et al., 2022) reported that detection of E. coli was significantly more common in samples from June to November. The drivers of this seasonality are not fully understood but could be related to higher temperatures promoting bacterial growth, higher levels of contaminated irrigation waters and/or a greater proportion of crops being grown in open fields rather than under cover.
All E. coli isolates from this survey were susceptible to all antibiotics tested in accordance to Assimilated Decision 2020/1729/EU. A previous UK survey in 2013-14 investigated the prevalence of AMR E. coli in fresh produce (lettuce, fresh herbs, fruits and vegetables) collected from leading supermarkets, hard discounter stores, convenience stores and local greengrocers (Randall et al., 2017). This study reported only one (a sample of carrots grown in the UK) of 400 fruit and vegetable samples with an E. coli level >10 CFU/g and no ESBL-producing nor carbapenem-resistant E. coli were detected, in line with the results presented here. A survey conducted in Belgium (2011-12) reported >100 CFU/g E. coli in 5% of 264 lettuce samples (Holvoet et al., 2013). Using disc diffusion and clinical interpretative criteria, resistance to ampicillin, amoxicillin-clavulanic acid or cephalothin was reported in over 10% of isolates recovered from lettuce leaves but none of the isolates showed resistance to cefotaxime. Resistance to streptomycin, sulphonamides, tetracycline and trimethoprim was also reported at lower levels (Holvoet et al., 2013). Similarly, a survey in the United Arab Emirates (2022-23) reported >100 CFU/g E. coli in 5.4% of 130 lettuce samples and in 26.5% of 400 salad leaves (Habib et al., 2023). In this survey, 145 isolates originating from lettuce and other salad leaves (spinach, rocket, herbs and arugula) were screened for antibiotic resistance using disc diffusion and clinical breakpoint values. The isolates exhibited the highest phenotypic resistance to ampicillin and tetracycline (≥ 20%), with 20 isolates exhibiting a multidrug-resistant phenotype (non-susceptible to three or more antimicrobial classes). None of the isolates were resistant to imipenem, and a limited proportion (< 10%) of the isolates showed resistance to clinically significant antibiotics such as ceftriaxone, cefotaxime, ciprofloxacin, and azithromycin (Habib et al., 2023).
The prevalence of Salmonella and STEC in lettuce has been low in previous surveys of lettuce and other salad products. McLauchlin (McLauchlin et al., 2022) did not recover any Salmonella or STEC from 201 salad leaves samples from UK retail and catering premises collected in 2020-21, although one STEC isolate was recovered from prepacked mixed salad in this survey. Similarly, a recent salad survey isolated three (0.1%) STEC from out of 2,495 salad ingredients, including one STEC from an iceberg lettuce, one from a sample of spinach and one from a mixed leaf salad (Kesby et al., 2026).
Between 2004 and 2012, Salmonella were isolated from seven out of 10,816 whole lettuce (romaine and iceberg) samples collected as part of the Microbiological Data Program survey of fresh produce at the USA, giving a prevalence of 0.06% (Reddy et al., 2016). The same study reported low prevalence of antibiotic resistant Salmonella in fresh fruit and vegetables, with only three out of 13 Salmonella isolates from lettuce showing any resistance. Zhang (Zhang et al., 2018) reported one and three Salmonella detections, none and one E. coli O157:H7 detections and none and five non-O157 STEC detections on 3,128 and 4,354 iceberg and romaine lettuce samples respectively, collected during 2010-2012 in the United States.
In the current survey of whole-head lettuce in the UK, no STEC or Salmonella were isolated, but traces of stx DNA (gene encoding for the Shiga toxin) were found in one sample of little gem lettuce; this could reflect detection of non-viable STEC cells, free stx-encoding bacteriophages and/or that bacteria other than STEC could interfere with the detection of low numbers of STEC (Kesby et al., 2024; Macori et al., 2020) . A recent survey detected stx DNA in 26 samples of salad components, but STEC was only isolated from three of them (Kesby et al., 2026).
The results of this survey have provided reassurance of the low microbiological risk to UK consumers from STEC, Salmonella and AMR bacteria in whole-head lettuce samples available at retail supermarket stores across the UK. However, the higher rates of E. coli and AMR E. coli contamination in catering premises and pre-packed products reported in previous surveys, along with the association of lettuce and other fresh produce with outbreaks of foodborne pathogens, highlight the need for continued monitoring of microbiological risks in lettuce and other fresh produce. We encourage growers and food business operators to continue to maintain robust Hazard Analysis and Critical Control Points (HACCP) plans and Good Agricultural Practice (GAP) to minimise microbiological contamination in the fields as well as cross-contamination and pathogen growth during storage and processing of whole-head lettuce.

_and_.png)
.png)
_and_.png)
.png)